The problem of prostate cancer detection and management
“Every year, more than a million men undergo painful needle biopsies for prostate cancer, and upward of 100,000 have radical prostatectomies, resulting in incontinence and impotence. But the shocking fact is that most of these men would never have died from this common form of cancer, which frequently grows so slowly that it never even leaves the prostate. How did we get to a point where so many unnecessary tests and surgeries are being done?”. This is a quote from the book published in the 2014, “The Great Prostate Hoax” by Professor Richard J. Ablin – the man who identified prostate specific antigen (PSA) and for decades fought against the misuse of his discovery and the “one size does not fit all” paradigm [1].
Being the second most common cancer in men, with constantly rising incidence, generally PCa is not highly lethal: according to recent National Cancer Institute SEER data, 98% of patients with newly diagnosed disease survive 5 years. Despite this encouraging data, such a high survival rate does not apply to patients in late stages of the disease and/or with poorly differentiated PCa of grade groups 4-5 according to the International Society of Urological Pathology (ISUP), with 5-year biochemical recurrence-free survival rates not exceeding 50.4% and 23.5%, accordingly. For the above-mentioned cohorts of patients, early and personalized detection of PCa is essential in order to grant maximally effective treatment. However, it is still unclear if PSA screening on the population level could bring more advantages than harm: according to a recent Cochrane review, based on the results of 4 available randomized controlled trials, no overall survival (OS) benefit was observed (RR: 1.00, 95% CI: 0.96-1.03), while screening was associated with such drawbacks as over-diagnosis and over-treatment [2,3]. The words of R. Ablin perfectly illustrate the current situation in the diagnostics of PCa, and particularly the inability of PSA as a tumour marker to efficiently detect this pathology (due to low specificity ~21% and sensitivity ~83%) as well as to stratify patients who require definitive treatment or only active surveillance. The situation is worsened by an enormously high rate of false-negative results from puncture biopsy in the diagnostics of PCa, which reaches 45%.
Genomics: molecular marker-based personalized approaches to prostate cancer
The urge to guide treatment tactics based on personal clinical risk factors has evolved in the era of human genome sequencing. Yet, unlike other malignancies such as breast cancer or glioblastoma, personalized approaches to managing PCa patients have not yet been developed. In recent years, a number of validated secondary oncomarkers, with improved specificity for PCa, appeared on the clinical scene and have already been included into American and European recommendations under the term “liquid biopsy”. Among them are such long non-coding urine-based mRNAs as Prostate Cancer Antigen 3 (PCA3) [4], HOXC6, DLX1 [5], and TMPRSS2-ERG (T2:ERG) [6], which are called on to help determine whether repeat puncture biopsy is needed after an initially negative biopsy. An example of a complex personalized approach to PCa risk calculation is incorporated into the Prostate Cancer Prevention Trial Risk Calculator patients’ clinical data, including levels of PSA, free PSA, and expression of PCA3 and T2:ERG. Nevertheless, clinical and cost-effectiveness, as well as efficiency in detection, of biochemical recurrence (BR) by means of the above-mentioned markers remains uncertain [2]. According to the promising results of several contemporary meta-analyses of transcriptomic data investigation, a spectrum microRNA (miR) has shown good diagnostic and predictive potential in the discrimination of PCa from benign prostatic hyperplasia/normal controls, being significantly upregulated (miR-18a, miR-21, miR-34a, miR-106b, miR-141, miR-182, miR-183, miR-200a/b, miR-301a, and miR-375) or downregulated (miR-1, miR-23b/27b, miR-30c, miR-99b, miR-139-5p, miR-152, miR-187, miR-204, miR-205, miR-224, miR-452, miR-505, and let-7c) [7,8]. Furthermore, Song et al. demonstrated that overexpression of miR-32 and underexpression of let-7c allowed differentiation of metastatic PCa from local/primary PCa, while expression profiles of 5 miRNAs (miR-21, miR-30c, miR-129, miR-145, let-7c, and miR-375) were associated with poor relapse-free survival or worse OS [7]. Such data illustrates the paramount role of miRNAs as regulators of PCa progression. However, none of those potential markers was externally validated, nor are there any substantial data about their value in a personalized clinical setting.
Recent studies have expanded the scope of the potential personalized approach to PCa screening even further: according to obtained data, men who inherit germline mutations in BRCA1, BRCA2, ATM, CHEK2, and MSH2/MSH6 genes are at increased risk of developing aggressive PCa. For men with a BRCA1/2 germline mutation, an individualized risk-based PCa screening approach was proposed by Cheng et al.: at age 40 years a measurement of baseline PSA and digital rectal exam, followed by, in the event of abnormal results, an MRI, biopsy, or further monitoring [9]. Future research will be critical for the improvement of screening tactics for men who are at the highest risk for aggressive PCa.
The issue of a personalized approach to the prognosis of the effectiveness of PCa treatment is another major clinical problem, due to significant divergence of oncologic post-treatment outcomes amongst patients stratified into the same risk category. Contemporary genome profile sequencing, and analysis of proteomic, transcriptomic, and metabolomic data have made it possible to define novel markers of prognosis and treatment response. Thus, Quero et al. found that measurement of miR-210 expression could be used as an endogenous marker of chronic hypoxia of PCa cells, which is directly connected with the radioresistance of a tumour [10]. Gong et al. demonstrated that in patients with miR-145 overexpression accompanied by underexpression of DNA repair genes regulated by miR-145, a good response to neoadjuvant radiotherapy of PCa was observed [11]. Moreover, Drake et al. developed a clinically relevant hierarchy of therapeutic kinase targets and involved pathways of patients with metastatic castration-resistant PCa, in order to personalize kinase signature (patient Cancer Hallmark Integrated Phospho Signatures; pCHIPS), aiming to lead clinical decisions, predict the most efficient drug combination, minimize treatment toxicity, and optimize precision-targeted treatment [12]. Overall, research in the given direction is only in the initial stages and currently no markers are validated nor approved for wide clinical implementation. Further investigation is required in order to overcome unmet needs in personalized prognostication of PCa treatment efficiency.
Radiomics: imaging markers in the management of prostate cancer
What will be the answer to the question of whether molecular markers alone can provide a meaningful and comprehensive approach to the personalized management of PCa patients? Undoubtedly, the answer will be negative, given the lack of information obtained through genetic markers regarding other, prognostically important visual features of the tumour, primarily stage, response to treatment, or the appearance of postoperative relapse. At the same time, how valuable is imaging of PCa alone, without liquid oncomarkers? As the mainstay of PCa imaging, multiparametric MRI (mpMRI) provides data, which is essential for all aspects of patient management - from primary diagnostics (according to the Prostate Imaging-Reporting and Data System; PI-RADS system, v2.1) to post-treatment monitoring. In contrast to CT and USG, mpMRI collects information not just about the spatial or textural parameters of the prostate lesion, but also about functional and pharmacokinetic aspects of PCa tissue. Currently, one of the most investigated MRI-based imaging markers of localized PCa is the apparent diffusion coefficient (ADC) measured from diffusion-weighted images (DWI), which represent the degree of restriction of Brownian motion of hydrogen molecules in cancer tissues [13]. According to recent meta-analyses, ADC values showed moderate accuracy in separating high-risk from low-risk PCa in accordance to ISUP grades (76.9% sensitivity, 77.0% specificity, AUC = 0.67) [14] and also allowed detection of extraprostatic extension of PCa with polled sensitivity and specificity 80.5% and 69.1%, respectively [15]. Notwithstanding, there are 2 major unsolved diagnostic problems inherent for mpMRI: the inability to reliably distinguish PCa from benign processes in PI-RADS 3 lesions (equivocal risk) and the predominant invisibility of aggressive cribriform/glomeruloid Gleason 4 patterns on MR-images [16]. In the context of advanced PCa, promising results were obtained by Zamboglou et al., demonstrating the feasibility of radiomic feature analysis of prostate-specific membrane antigen PET for discrimination of intraprostatic tumour and non-invasive characterization of ISUP grade as well as for pelvic lymph node status characterization [17]. A spectrum of other potential functional (diffusion kurtosis, blood oxygen level-dependent, diffusion tensor imaging) and perfusion imaging markers (Ktrans, Kep, Ve, arterial spin labelling) still have not received wide clinical usage and require more profound inquiry.
Radiogenomic analysis in prostate cancer patients: possibilities, limitations,
and perspectives
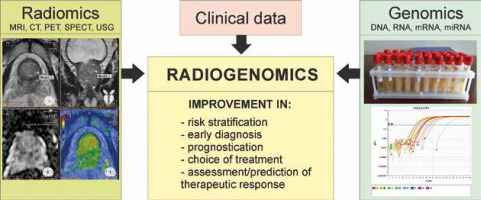
Radiogenomics (i.e. imaging genomics) is a relatively new term, used to refer to the study of genetic variation associated with imaging features of the tumour (Figure 1). Several recent works have proven that by combining individual tumour features, such as hypoxia status, genomic/transcriptomic and radiomic profiling (radiologic-molecular correlation), with traditional staging procedures in order to personalize treatment for PCa, an improved prognostication of PCa can be achieved and overtreatment of “clinically insignificant” (i.e. indolent) cancer can be avoided [18,19]. In 2015 Renard-Penna et al. identified prognostic biomarkers of PCa using radiogenomics analysis by integration of the gene expression using the cell cycle progression score and MRI data. It was found that the combined measure of maximal lesion diameter < 10 mm and ADC > 0.80 × 10-3 mm2/s identified exclusively tumours harbouring primary Gleason grade 3. However, the cell cycle progression score did not completely match the mpMRI data: 7 of those lesions demonstrated a molecular pattern of clinically significant lethal prostate cancer (cell cycle progression score > 0) [20]. A few years ago, Stoyanova et al. combined mpMRI data with gene expression analysis and 2-way hierarchical clustering; as a result 49 radiomic features of PCa correlated with 3 gene signatures associated with adverse outcome (Gleason 8-9 disease). Genes that demonstrated very high positive correlation (≥ 0.9) to radiomic MR-parameters were TRPM8, DPP4, and GCNT1 [21]. McCann et al. revealed a weak but significant association between the quantitative perfusion parameter of the dynamic contrast-enhanced MRI – k(ep) with PTEN expression (r = –0.35, p = 0.02) [22]. Consequently, Jamshidi et al. amalgamated mpMRI data and whole-exome spatial multiregional characterization of PCa microenvironments. As a result, 77 mutations involving 29 cancer-associated genes across PCa tissue samples were identified, which allowed, by means of hierarchical clustering, the separation of high-grade lesions from the normal tissues. The genes with the largest weightings in distinguishing patients with Gleason scores of 3+4 versus 4+5 were KLK2, KRAS, SPINK1, BRCA1, and BCR. There was no significant difference in mutation load in cancer-associated genes between regions that were proven to be normal via histopathological analysis (34 mutations per sample), mildly suspicious via multiparametric MR imaging, intermediately suspicious (31 mutations per sample), and high-grade cancer (33 mutations per sample) (p = 0.30). As a result, the continuum of mutations in PCa and normal prostate tissues according to whole-exome radiogenomic analysis and spatial multiparametric MRI of prostate was demonstrated [23]. In a recently published work by Fischer et al.,significant efforts were made to decode the underlying PCa progression molecular mechanisms by means of a radiogenomic approach. The authors performed correlative analysis of MRI data (segmentation, extraction of aggressiveness-related features, histogram of volume intensity and texture features) and gene/miRNA expression profiles for T2c and T3b PCa stages. It was revealed that biomarkers such as ANPEP, miR-217, miR-592, and miR-6715b may drive tumour upstaging, and it allowed the PCa stages to be distinguished; they also correlated highly (r = 0.75) with corresponding aggressiveness-related radiomic MR-features [24]. To address the issue of the invisibility of some PCas on mpMRI, Houlahan et al. profiled the genomes and transcriptomes of patients with ISUP grade 2 tumours: mpMRI-invisible (PI-RADS < 3) and mpMRI-visible (PI-RADS 5) lesions. For mpMRI-visible cancers, genomes with amplified quantity of mutations, a higher prevalence of cribriform pattern of growth, and a spectrum of 102 altered transcripts (upregulation of noncoding RNAs such as SCHLAP1) were habitual. Also, multiple small nucleolar RNA hallmarks were identified in order to discriminate visible from invisible PCs [25].
Conclusions
To summarize, being an attractive research topic, the radiogenomics of PCa currently is not a comprehensively investigated area of oncourology. According to preliminary research findings conducted in this field, the combination of genomics and radiomics (and presumably metabolomics, proteomics, and transcriptomics) as integrative parts of precision medicine in the future has the potential to become the foundation for a personalized approach to the management of PCa. However, there are a number of hindrances to achieving this goal, such as relatively small numbers of patients included in current studies, a lack of available large randomized controlled trials, the need to use complex integrated methods of big data analysis, the comparatively high cost of genomic profiling and imaging methods, and the question of whether, before we include any potential genomic or transcriptomic marker into radiogenomic analysis, it should first be validated in order to prove its separate clinical value. If so, it greatly and significantly shifts the horizon of the actual use of radiogenomics in clinical practice, owing to the need for a huge body of future research.